|
Importing Data from ScanArray Files
Overview
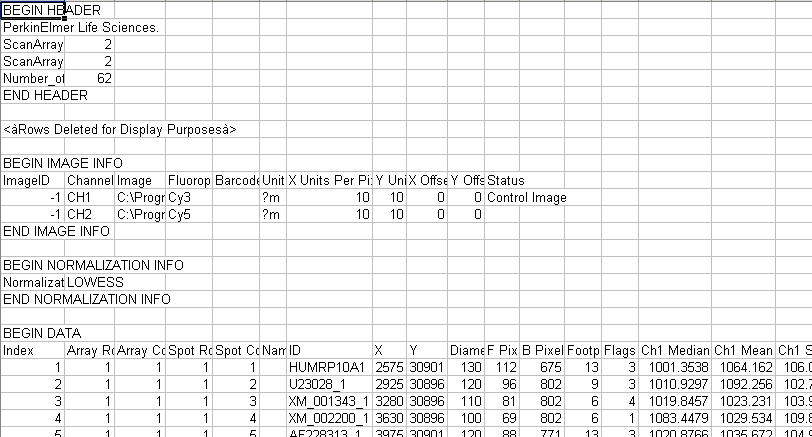
The data files must be in the Perkin-Elmer ScanArray file format.
Sample Order
The sample order of imported datasets is determined by the order of the source sample data files listed in the Import Data dialog.
|
Template |
Result of Import |
|
ScanArray |
Multiple files are processed into a single dataset. |
|
ScanArray Merge Replicates |
Multiple files are processed into a single dataset. |
|
ScanArray Ch1/Ch2 |
Multiple files are processed into a single ratio dataset (treatment/control). |
|
ScanArray Ch2/Ch1 |
Multiple files are processed into a single ratio dataset (treatment/control). |
Import Process for ScanArray and ScanArray Merge Replicates
The file headers are discarded.
Gene identifier information is retrieved from the Name column of the first file and is stored as a GenBank Identifier.
Gene expression data is retrieved from the Ch2 Ratio of Medians column of each file in the order they are placed in the Import Data dialog.
Import Process for ScanArray Ch1/Ch2
The file headers are discarded.
The RatioFormulation field is ignored.
Gene identifier information is retrieved from the Name column of the first file and is stored as a GenBank Identifier.
The control (Ch2) expression data is calculated by subtracting the Ch2 B Median column from the Ch2 Median column.
The treatment (Ch1) expression data is calculated by subtracting the Ch1 B Median column from the Ch1 Median column.
Import Process for ScanArray Ch2/Ch1
The file headers are discarded.
The RatioFormulation field is ignored.
Gene identifier information is retrieved from the Name column of the first file and is stored as a GenBank Identifier.
The control (Ch1) expression data is calculated by subtracting the Ch1 B Median column from the Ch1 Median column.
The treatment (Ch2) expression data is calculated by subtracting the Ch2 B Median column from the Ch2 Median column.
Related Topics:
Selecting a Template for Data Import
Importing Multiple Files With One Sample Each